Record building workflow#
NexusLIMS automatically builds XML records from electron microscopy data by combining session information from the database with metadata extracted from data files. This page explains the workflow from session detection to record upload. See the activity diagram at the bottom for a visual overview.
Understanding these details isn’t required for basic operation, but this documentation may help you troubleshoot issues, customize behavior, and understand what the system does behind the scenes. Links to API reference documentation are provided throughout.
General Approach#
Since instruments cannot communicate directly with NexusLIMS, the system uses periodic
polling (via systemd or cron) to detect sessions requiring record generation.
Execution: Run nexuslims build-records to invoke
process_new_records(). See the
CLI reference for all command-line options including date filtering,
dry-run mode, and verbosity controls. The command:
Queries NEMO APIs for recent usage events
Finds database sessions awaiting record generation
Searches the centralized file system for matching files
Extracts metadata and groups files into
Acquisition ActivitiesBuilds and validates XML records against the Nexus Microscopy Schema
Uploads valid records to the CDCS instance (and any other export destinations configured)
Error handling: Failed builds are logged to the database with ERROR status,
triggering email notifications.
Finding New Sessions#
process_new_records() discovers sessions
requiring record generation through two steps:
Harvest NEMO events:
add_all_usage_events_to_db()queries configured NEMO APIs for recent usage events (default: 7 days lookback) and adds them to the session log database.Query pending sessions:
get_sessions_to_build()finds sessions withTO_BE_BUILTstatus:SELECT (session_identifier, instrument, timestamp, event_type, user) FROM session_log WHERE record_status == 'TO_BE_BUILT';
Results are converted to SessionLog objects,
then paired into Session objects by matching
START and END logs with the same session_identifier.
Session Attributes#
Each Session contains:
session_identifier(str): Session ID ( resolvable URL for NEMO events)instrument(Instrument): Associated instrument objectdt_from(datetime): Session start timedt_to(datetime): Session end timeuser(str): Username (read from NEMO usage event)
Sessions are processed sequentially by the record builder.
Building a Single Record#
For each Session, the record builder
executes these steps:
Process Overview#
(link) Initiate build: Execute
build_record()(link) Fetch usage event/reservation data: Query harvesters (e.g.,
nemo)(link) Find files: Search for parseable files within session timespan
If none found → mark as
NO_FILES_FOUNDand skip to next session
(link) Cluster activities: Group files into
AcquisitionActivityobjects using temporal analysis(link) Extract metadata: Parse each file’s metadata and generate preview images
(link) Organize metadata: Separate common metadata (setup parameters) from file-specific values
(link) Validate: Check XML against Nexus Microscopy Schema
(link) Upload: Push valid records to CDCS/other export destinations and update database
1. Initiating the Build#
Before building each record:
insert_record_generation_event()logs the build attempt to the database for audit purposesbuild_record()is invoked with the session’s instrument and timestamp informationBasic XML header is written before querying the reservation system
2. Querying the Reservation/Tool Usage Calendars#
Instrument reservation and usage tracking systems (like NEMO) provide experiment metadata
(user, sample, motivation, etc.) for the record’s <summary> element.
The record builder queries the harvester specified in the instrument’s database entry
(typically nemo). Each harvester implements
res_event_from_session(), which returns a
ReservationEvent object representing
the reservation with maximum time overlap (or a generic event if none found).
Added in version 2.3.0: While the class is named ReservationEvent for historical reasons, it can contain
metadata harvested from either Reservations or Usage Events (actual instrument
usage sessions). This capability prioritizes Usage Event data since it
reflects what was actually performed, falling back to Reservation data when usage
information is unavailable.
This adapter layer provides uniform reservation data regardless of the source system. The event is then serialized to schema-compliant XML.
Three-Tier Fallback Strategy for NEMO#
For NEMO-based systems, the harvester uses a three-tier fallback strategy to obtain experiment metadata, prioritizing the most recent and accurate information:
Priority 1: Post-Run Questions (run_data)
The highest priority source is the run_data field from NEMO usage events. This field
is populated when users answer questions at the end of their instrument session
(when they “enable” the tool to end their usage). Since these questions are answered
after the experiment completes, they represent the most accurate description of what
was actually performed.
Priority 2: Pre-Run Questions (pre_run_data)
If run_data is empty or invalid, the harvester falls back to the pre_run_data field.
This field is populated when users answer questions at the start of their instrument
session (when they initially “enable” the tool). While potentially less accurate
than post-run data (since the experiment plan may change during execution),
this provides valuable metadata when users don’t complete post-run questions.
Priority 3: Reservation Matching
If neither usage event question data source is available, the harvester falls back to traditional reservation matching. The system finds the reservation with maximum time overlap with the usage event and extracts metadata from the reservation’s question data. This is the least reliable method since reservations may be created well in advance and might not reflect the actual experiment performed.
Question Data Structure
All three sources (run_data, pre_run_data, and reservation question_data) use
identical JSON structure, enabling consistent metadata extraction. Both usage event
fields are JSON-encoded strings that are parsed and validated during harvesting.
See res_event_from_session() documentation for
the required question structure and examples.
Data Consent Validation#
All metadata sources require affirmative user consent via a data_consent question.
The harvester validates consent for each source in priority order:
Missing consent field: Falls back to next priority level or raises
NoDataConsentErrorDeclined consent (“Disagree”, “No”, etc.): Falls back to next priority level or raises
NoDataConsentErrorInvalid JSON or missing question data: Falls back to next priority level
If all three tiers fail (no valid question data with consent), the session is marked
NO_CONSENT or NO_RESERVATION in the database and no record is generated.
Graceful Degradation#
The three-tier system provides graceful degradation:
Empty strings, malformed JSON, or missing fields in higher-priority sources automatically trigger fallback to the next tier
Each tier is validated independently for data completeness and consent
Logging tracks which data source was ultimately used for each session
The system prioritizes data quality over data availability
This approach maximizes record generation success while preferring the most accurate metadata source available for each session.
3. Identifying Files to Include#
Record metadata is primarily extracted from session files. The
Session provides the instrument,
timespan, and identifier. Instrument configuration (persistent ID, location,
file storage path relative to NX_INSTRUMENT_DATA_PATH)
is read from the database instruments table, centralizing configuration management.
File search strategy:
gnu_find_files_by_mtime() uses GNU
find to find files modified
within the session timespan.
No files found: Sessions without matching files (accidental session start, no data generated)
are marked NO_FILES_FOUND and the builder moves to the next session.
(go to top)
4. Separating Acquisition Activities#
Files are grouped into logical AcquisitionActivity
objects to approximate conceptual experiment boundaries using statistical analysis of
file creation times.
Added in version 2.3.0: The NX_CLUSTERING_SENSITIVITY configuration variable allows you to adjust how files are clustered into activities. Higher values (e.g., 2.0) create more activities by detecting smaller time gaps, while lower values (e.g., 0.5) create fewer activities by requiring larger gaps. Set to 0 to disable clustering entirely and group all files into a single activity.
The figure below illustrates the clustering process for an EELS spectrum image experiment. Panel (a) shows creation time differences revealing 13 distinct groups. Panel (b) histograms time gaps between consecutive files—small gaps indicate related files, large gaps mark activity boundaries.
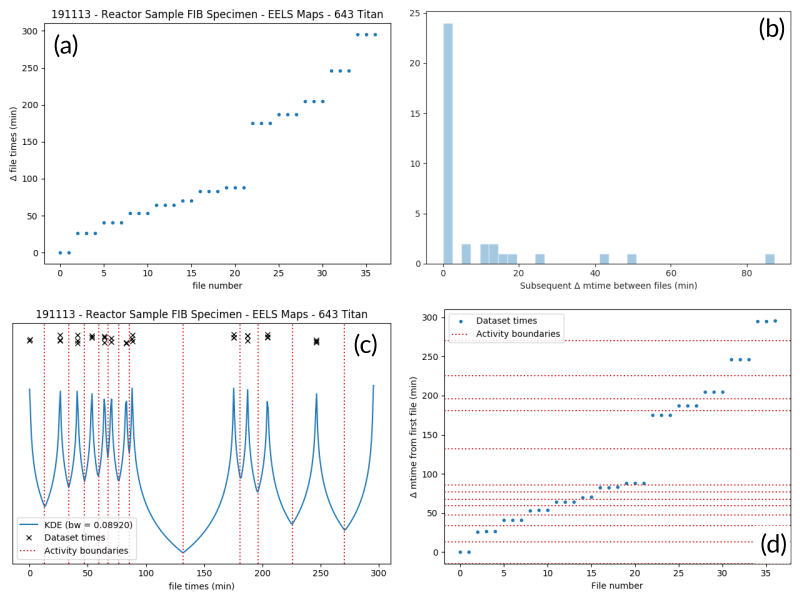
An example of determining the
AcquisitionActivity time boundaries
for a group of files collected during an experiment. See the surrounding
text for a full explanation of these plots.#
Clustering algorithm: cluster_filelist_mtimes()
uses Kernel Density Estimation
(KDE) to detect temporal gaps. The KDE peaks where files cluster in time and minimizes
at gaps. Grid search cross-validation optimizes the KDE bandwidth for each session.
Local minima become activity boundaries (panel c).
Panel (d) overlays boundaries on the original time plot, successfully identifying all 13 groups as a human would. This approach generalizes well across file types and experiment patterns.
5. Parsing Individual Files’ Metadata#
parse_metadata() dispatches to format-specific extractors
in nexusLIMS.extractors. Each extractor returns a dict with native
metadata plus a top-level 'nx_meta' key containing normalized metadata for the XML record.
Metadata Validation (v2.2.0+):
Pydantic schemas (nexusLIMS.schemas.metadata) validate metadata quality:
Schema selection by
DatasetType(ImageMetadata,SpectrumMetadata, etc.)Required fields:
creation_time(with timezone),data_type,dataset_typeNormalization: EM Glossary terms (where available), unit standardization (voltages → kV, distances → nm/mm)
Type safety: Invalid types, missing timezones, incorrect enums trigger logged errors
This evolving system provides foundational standardization. See Internal Metadata Schema System for details.
XML serialization: Nested 'nx_meta' structures are flattened (space-separated keys).
Most key-value pairs become Setup Parameters or Dataset Metadata in the CDCS front-end.
Special metadata keys
Reserved 'nx_meta' keys with special handling:
DatasetType: Controlled vocabulary mapped to<dataset @type>(Image, Spectrum, etc.)Data Type: Human-readable descriptor (TEM_Imaging,SEM_EDS,STEM_EELS) for activity contents summaryCreation Time: ISO-8601 timestamp displayed in dataset tableExtraction Details: Metadata extraction provenance informationwarnings: List of unreliable metadata keys (front-end displays warnings)
See Internal Metadata Schema System for complete metadata schema documentation.
Extractor implementation: Extractors leverage HyperSpy for most formats. Unsupported formats require manual binary decoding and custom extractor/preview generator implementation.
Output artifacts: parse_metadata() generates (saved to
NX_DATA_PATH, mirroring NX_INSTRUMENT_DATA_PATH structure):
JSON metadata file: Full metadata in text format (linked from XML record)
PNG preview image: Generated via
nexusLIMS.extractors.plugins.preview_generators(HyperSpy-based for complex formats, simple downsampling for regular image files)
Metadata and preview paths are stored at the
AcquisitionActivity level.
6. Determining Setup Parameters#
After file processing, metadata is organized hierarchically:
Common values (identical across all activity files) →
<AcquisitionActivity>Setup ParametersUnique values (varying per file) →
<dataset>metadata
store_setup_params() identifies
common metadata by comparing values across files.
store_unique_metadata() then
stores only file-specific values at the dataset level.
This organization helps users distinguish constant experimental conditions from per-file variations.
7. Validating the Built Records#
Each completed AcquisitionActivity is converted
to XML via as_xml(). Once all
activities are assembled, validate_record()
validates the complete record against the NexusLIMS schema.
Validation failure: Session marked ERROR in database for investigation.
Validation success: Record written to NX_RECORDS_PATH (or NX_DATA_PATH subdirectory if unspecified) before uploading to configured exporters.
The builder then processes the next session, repeating until all are complete.
8. Uploading Completed Records and Updating Database#
After all sessions are processed, export_records()
uploads generated XML records to configured export destinations.
Upload process:
CDCS re-validates each record against the NexusLIMS schema
Records are assigned to the
Global Public Workspace(viewable without login)Database is updated with upload status
Note
Future versions will implement single-sign-on, assigning records to their creating users instead of the public workspace.
The complete workflow repeats periodically (see General Approach) to process new sessions as they occur.
Record Generation Diagram#
The following diagram illustrates the logic (described above) that is used to
generate Experiment records and upload them to the NexusLIMS CDCS instance.
The diagram shows the flow through different modules/sub-packages within the
nexusLIMS package. Decision points determine the path through the workflow,
and database updates occur at key stages to track session status.
Tip
💡 Interactive Diagram: You can zoom in/out (scroll or pinch) and pan (click and drag) to explore different parts of the workflow in detail.
%%{init: {'theme': 'base', 'themeVariables': { 'primaryColor': '#eee', 'primaryTextColor': '#000', 'primaryBorderColor': '#333', 'lineColor': '#333', 'secondaryColor': '#eeeeff', 'tertiaryColor': '#eeffee', 'noteBkgColor': '#fff9e6', 'noteBorderColor': '#d4a017'}}}%%
flowchart TD
Start([Start]) --> ProcessNewSessions["record_builder.process_new_sessions()"]
ProcessNewSessions --> AddUsageEvents["harvesters.nemo.add_all_usage_events_to_db()"]
AddUsageEvents --> GetSessions["session_handler.get_sessions_to_build()"]
GetSessions --> CheckSessions{New sessions<br/>found?}
CheckSessions -->|No| End1([End])
CheckSessions -->|Yes| ExtractSession["Extract single Session"]
ExtractSession --> ReturnSession["Return session_handler.Session object"]
ReturnSession --> InsertEvent["Insert RECORD_GENERATION event<br/>in database"]
InsertEvent --> BuildRecord["record_builder.build_record(Session)"]
BuildRecord --> CreateXML["Create new .xml document<br/>for this Experiment"]
CreateXML --> FindReservation["harvesters.XXXX.res_event_from_session(s)"]
FindReservation --> CheckReservation{ReservationEvent<br/>found?}
CheckReservation -->|Yes| AddDetailedInfo["Add detailed event info<br/>to record summary XML"]
CheckReservation -->|No| AddBasicInfo["Add basic information<br/>(instrument and date)"]
CheckReservation -->|No consent| UpdateNoConsent["Update Session status<br/>as NO_CONSENT"]
CheckReservation -->|No reservation| UpdateNoReservation["Update Session status<br/>as NO_RESERVATION"]
AddDetailedInfo --> IdentifyFiles
AddBasicInfo --> IdentifyFiles["Identify files created on this<br/>instrument within timespan"]
IdentifyFiles --> CheckFiles{Files found?}
CheckFiles -->|No| UpdateNoFiles["Update Session status<br/>as NO_FILES_FOUND"]
CheckFiles -->|Yes| ClusterFiles["Separate files into activities<br/>by clustering file creation times"]
ClusterFiles --> ProcessActivity["Build one AcquisitionActivity"]
ProcessActivity --> ParseMetadata["Parse metadata for all files"]
ParseMetadata --> ExtractLoop["extractors.parse_metadata(filename)"]
ExtractLoop --> ExtractMetadata["Extract metadata from file"]
ExtractMetadata --> DetermineType["Determine dataset type"]
DetermineType --> ValidateMetadata["Validate metadata using<br/>Pydantic schemas"]
ValidateMetadata --> GeneratePreview["Generate .png preview image"]
GeneratePreview --> SaveMetadata["Save metadata as .json"]
SaveMetadata --> CheckMoreFiles{More files<br/>to process?}
CheckMoreFiles -->|Yes| ExtractLoop
CheckMoreFiles -->|No| AssignMetadata["Assign common metadata as<br/>Setup parameters and<br/>unique metadata to datasets"]
AssignMetadata --> AddActivity["Add AcquisitionActivity<br/>to .xml record"]
AddActivity --> CheckMoreActivities{More activities<br/>to process?}
CheckMoreActivities -->|Yes| ProcessActivity
CheckMoreActivities -->|No| ValidateRecord{Validate<br/>record?}
ValidateRecord -->|Valid| WriteXML["Write XML record<br/>to NexusLIMS folder"]
ValidateRecord -->|Invalid| UpdateError["Update Session status<br/>as ERROR"]
WriteXML --> UpdateCompleted["Update Session status<br/>as COMPLETED"]
UpdateCompleted --> VerifyUpdate["Verify Session was updated<br/>in database"]
UpdateError --> VerifyUpdate
UpdateNoFiles --> VerifyUpdate
UpdateNoConsent --> VerifyUpdate
UpdateNoReservation --> VerifyUpdate
VerifyUpdate --> CheckMoreSessions{More sessions<br/>to process?}
CheckMoreSessions -->|Yes| ExtractSession
CheckMoreSessions -->|No| CheckRecordsCreated{Any records<br/>created?}
CheckRecordsCreated -->|No| End2([End])
CheckRecordsCreated -->|Yes| UploadRecord["Upload single .xml record"]
UploadRecord --> AssignWorkspace["Assign to public<br/>CDCS workspace"]
AssignWorkspace --> CheckMoreRecords{More records<br/>to upload?}
CheckMoreRecords -->|Yes| UploadRecord
CheckMoreRecords -->|No| End3([End])
%% Styling
classDef recordBuilder fill:#eee,stroke:#333,stroke-width:2px
classDef sessionHandler fill:#eeeeff,stroke:#333,stroke-width:2px
classDef extractors fill:#eeffee,stroke:#333,stroke-width:2px
classDef cdcs fill:#ffefe1,stroke:#333,stroke-width:2px
classDef dbUpdate fill:#ffcc99,stroke:#333,stroke-width:3px
class ProcessNewSessions,BuildRecord,CreateXML,IdentifyFiles,ClusterFiles,ProcessActivity,AssignMetadata,AddActivity,ValidateRecord,WriteXML recordBuilder
class AddUsageEvents,GetSessions,ReturnSession,InsertEvent,VerifyUpdate sessionHandler
class ParseMetadata,ExtractLoop,ExtractMetadata,DetermineType,ValidateMetadata,GeneratePreview,SaveMetadata extractors
class UploadRecord,AssignWorkspace cdcs
class UpdateCompleted,UpdateError,UpdateNoFiles,UpdateNoConsent,UpdateNoReservation dbUpdate
Diagram Key:
Module |
Light Mode |
Dark Mode |
|---|---|---|
Record Builder |
Gray boxes |
Dark gray boxes |
Session Handler |
Light blue boxes |
Dark yellow boxes |
Metadata Extractors |
Light green boxes |
Red boxes |
CDCS Connector |
Peach boxes |
Slate gray boxes |
Database Updates |
Orange boxes |
Light blue boxes |

